Apr 26, 2022 · 10 min read
5 Ways Single-Cell Biology Is Advancing Our Understanding of Disease
New methods come along every so often that profoundly change biomedical research and lead to dramatic improvements in human health. One example is DNA sequencing, which gave us the ability to read the human genome at scale. The technology has gotten so fast and inexpensive that it is now routinely used to diagnose and treat disease.
Building on those advances, researchers are also able to analyze the DNA and RNA in individual cells. Over the past decade, a technique known as single-cell sequencing has spurred research on variation in cells — the basic unit of life — and how they function. The field, known as single-cell biology, is poised to give us a more precise understanding of disease. Researchers are still asking many of the same fundamental questions. What has changed is the tools they use, and the scale and ease with which those new tools can be applied.
At the Chan Zuckerberg Initiative, Cell Science, (formerly the Single-Cell Biology program), is one of our flagship science programs. For the past five years, we have been supporting researchers assembling the Human Cell Atlas — an international effort to map all the cell types in the human body as a resource for better understanding health and disease. We have also been funding the development of new technologies that make it easier for researchers to conduct single-cell research, share their results and make potentially life-changing discoveries.
Single-cell sequencing is now coming of age. The field is no longer primarily driven by technology developers. It has become a much broader ecosystem, in which scientists and clinicians are generating key insights into biology and translating those to the clinic. Here are just a few exciting examples.
1. “Aging clocks” could help us counter aging and chronic disease.
As the world’s population ages, researchers are racing to understand healthy aging and how aging gives rise to chronic conditions such as dementia and cancer. Age after all is the greatest risk factor for most diseases. Among them is Anne Brunet, a geneticist at Stanford University.
“Understanding aging could provide new avenues to understand and counter many diseases at once,” Anne says.
But studying aging presents a problem. Unlike other biological processes such as breathing or sleeping, aging is hard to quantify. Sure, we use chronological age to mark the passage of time, but it doesn’t always mirror our biological age. And chronological age may mean something very different for cell types that have very long or very short lifespans. Without a way to measure aging, it’s difficult to study the age-related changes in our vital organs, cells and tissues, and the factors that slow down or speed up the aging process, such as inflammation.
Fortunately, Anne and her team have come up with a solution. They recently developed a set of “aging clocks” for different cell types in a regenerative region of the mouse brain. The clocks are mathematical models trained on data showing which genes are active in single cells collected from animals of different ages. When Anne put the aging clocks to the test, she found that they accurately predicted the cells’ chronological ages.
But she also wondered, could aging clocks help identify strategies to slow or counter aging? Anne and her collaborators tested how individual cell types respond to rejuvenation strategies — specifically exercise and blood transfusions — in a fascinating set of experiments. Remarkably, the clocks confirmed that both exercise and blood from young mice can turn back time in cells, changing the older cells’ gene activity to more closely resemble activity patterns in younger cells. The results also show that aging clocks could be used to assess rejuvenation in the brain and strategies to combat age-related diseases.
2. Massive lung cell atlas reveals the effects of smoking, infection and disease.
Just as the COVID-19 pandemic reminded us how precious our lungs are, researchers completed the largest and most detailed map of the respiratory system. The Human Lung Cell Atlas (HLCA) is an integrated atlas describing more than 2 million cells collected along the airway and the cells’ relationship to one another. This wealth of information will help researchers better understand how infections and diseases such as pulmonary fibrosis, asthma and cancer affect the cells and tissues of the lungs. Researchers have already used some of the HLCA data to identify the cell types most susceptible to SARS-CoV-2 infection and uncover the damaging effects of immune cells in asthma.
The power of the HCLA lies in its numbers. It integrates data collected worldwide from more than 400 people of diverse ancestries and demographics. Integrating the data from many different samples was a technical feat with a major payoff: The massive dataset made it possible to study the effects of variables like ethnicity, age, sex and smoking status on lung cells in health and disease.
“A comprehensive organ atlas should capture both cellular and individual diversity,” wrote Lisa Sikkema, the first author on a new preprint describing the HCLA, on Twitter. “It’s the first time the effects of these demographics on so many lung cell types could be modeled!”
Data from the HCLA is now available to patients, scientists, medical professionals and other experts who can explore it using Chan Zuckerberg CELL by GENE (CZ CELLxGENE), an open-source cell visualization and analysis tool developed by CZI.
3. Cells of the Fallopian Tube Linked to Cancer and Infertility

A new effort to characterize the fallopian tube is expanding our knowledge of reproductive health and our ability to treat infertility and reproductive diseases. The project is a collaboration between University of Michigan geneticists Sue Hammoud and Jun Li, clinician Erica Marsh and biomedical engineer Ariella Shikanov. They are working toward a human cell atlas of the female reproductive system.
The team used single-cell RNA sequencing to characterize the diversity of cell types in the fallopian tube — the site of fertilization. They identified 12 major cell types in healthy fallopian tissue, including several new types of epithelial cells with finger-like projections that help egg cells travel from the ovary to the uterus. They also found two types of progenitor cells, which can replenish or repair tissue.
“This really is a basecamp to launch future studies,” Jun says.
Future research could focus on understudied conditions such as infertility, endometriosis and ovarian cancer. In fact, initial findings from the fallopian cell atlas suggest the cellular roots of ovarian cancer may be in the fallopian tube. The researchers identified transition cells that could become cancerous under the right conditions.
4. An Autoimmune Disease Divulges Some of Its Secrets
Systemic lupus erythematosus, generally known as lupus, is an enigma. It’s an autoimmune disease that arises when the body’s defense system mistakenly attacks its own cells. But lupus is complicated to diagnose and treat because it can affect any number of organs. It is also unclear why lupus disproportionately affects women and individuals of Asian, African and Hispanic ancestry.
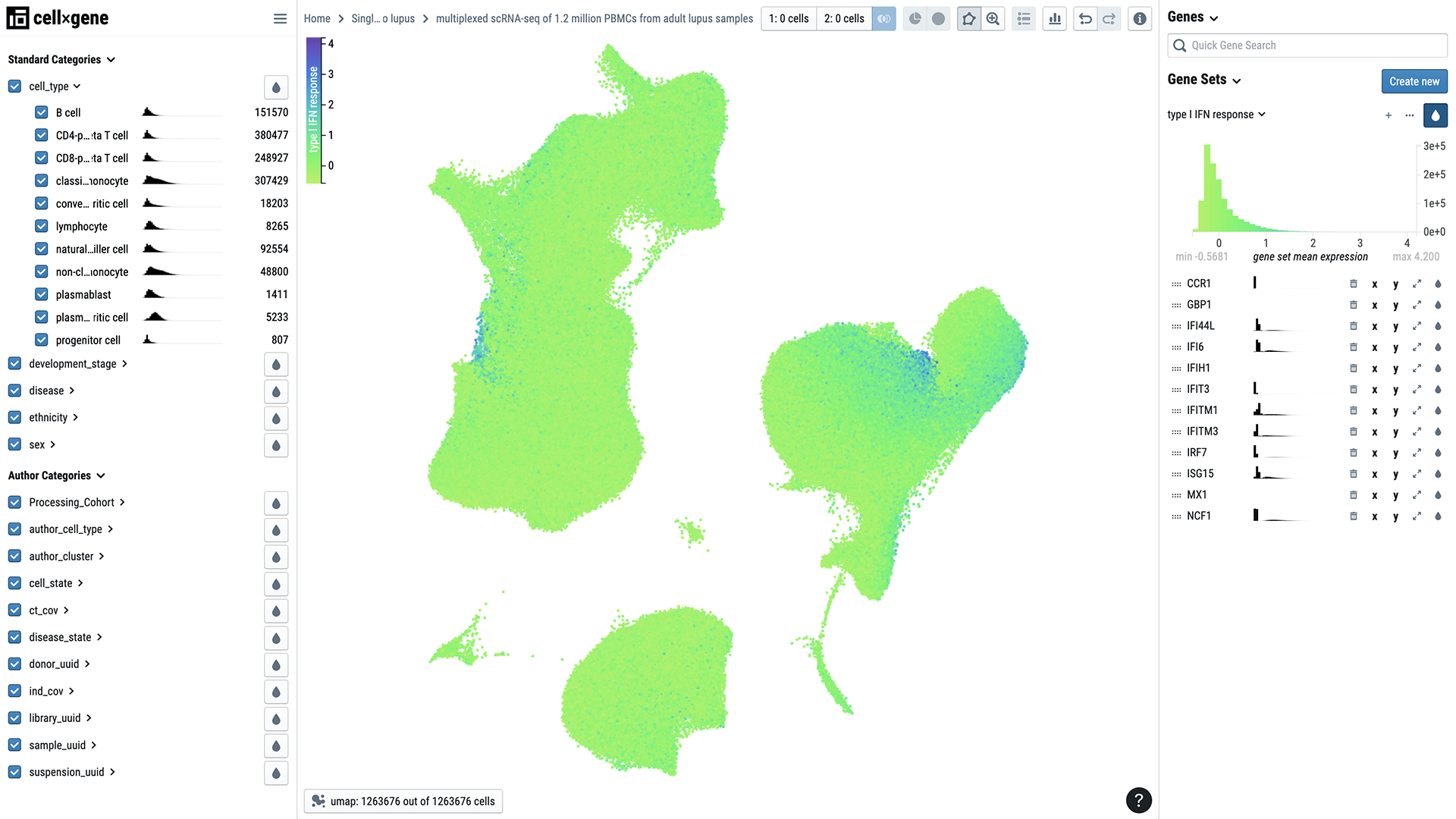
Single-cell biology is shedding light on some of these mysteries. Researchers recently characterized more than 1 million immune cells from 250 people, more than half of whom had lupus. The cells were primarily from people with European and Asian ancestry.
Until now, describing differences in so many cells from so many individuals has been a challenge. But the research team, led by biologist Jimmie Ye at the University of California, San Francisco, developed a new method to enable single-cell profiling of large populations.
The method is helping Jimmie and his collaborators uncover how differences in sex and ethnicity affect the mix of immune cells present in people with the disease. For example, they found that a certain population of immune cells is either diminished or expanded in some people with lupus. By probing these changes, researchers and clinicians may be able to figure out why some people are more susceptible to or have a more severe form of the disease than others.
The data, part of Human Cell Atlas, is being shared with researchers around the world via CZ CELLxGENE. It could spur the development of new computational methods for analyzing such a large number of samples and lead to new insights about immune disease, Jimmie says.
“We made the data available in CZ CELLxGENE so that other investigators could start analyzing it [as soon as possible],” he says.
5. Organoids Mirror Human Development and Help Engineer Cell Therapies

Barbara Treutlein’s research reflects her belief that “every cell has a story to tell.” The biologist, whose lab is based at the Swiss Federal Institute of Technology in Zürich, Switzerland, is working to uncover those cellular stories using tiny, three-dimensional tissue cultures grown from stem cells. These 3D lab-grown tissues, known as organoids, allow scientists to study how human organs develop and how they are changed by disease.
Barbara and her collaborators, like geneticist Gray Camp (University of Basel) and biologist Jason Spence (University of Michigan), want to understand to what degree organoids mimc complex tissues and how to create better organoids for research and therapeutic purposes. After all, the better the organoid, the more valuable it is for studying how human cells, tissues and organs develop and for engineering cell therapies.
Also read: How Lab-Made Stem Cells Are Transforming Research
Recently, Barbara, Gray and Jason produced a single-cell atlas of multiple organs derived from the endoderm (one of the three distinct cell layers in an embryo), including the intestine. They used the atlas to compare human intestinal cells to those cultured in the lab, and asked questions such as: Are the cell types the same? Do the cells mature in similar ways? And do they perform the same functions? They found that most, but not all, of the intestinal organoid cells mapped to intestinal counterparts in the atlas. Using their results, they are working to improve the organoids and explore aspects of human development that couldn’t be studied without them.