Essential Open Source Software for Science
CZI’s Essential Open Source Software for Science program supports software maintenance, growth, development, and community engagement for open source tools critical to science.
Showing 195 results
To empower African scientists to develop 3D Slicer AI-based extensions for health priorities and enable their deployment to global communities through customized AI models for multilingual translation. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Sonia Pujol (Brigham and Women's Hospital, Harvard Medical School)
To empower the biomedical research community in Latin America by localizing 3D Slicer to Spanish and Portuguese, improving tutorial localization infrastructure, and holding outreach events.

Sonia Pujol (Brigham and Women’s Hospital, Harvard Medical School)
To increase the accessibility of the 3D Slicer open source platform for biomedical research to clinicians and scientists in non-English speaking countries.

Sonia Pujol (Brigham and Women's Hospital, Harvard Medical School)
To develop scvi-tools 2.0: an accessible and scalable environment for probabilistic modeling for single-cell genomics that enables the next generation of massive-scale studies.
Nir Yosef (Weizmann Institute of Science)
To bring the combined power of scikit-image and Dash to a larger number of scientists thanks to increased execution speed, interactive image annotation and processing, and outstanding documentation targeting life sciences practitioners.


Emmanuelle Gouillart (Plotly Technologies, Inc.)
The project will improve the SciPy library's statistics functionality to better serve biomedical research and downstream projects. In addition, an outreach component will engage female students, inspiring them to participate in open source code development.

Warren Weckesser (University of California, Berkeley; NumFOCUS)
Matt Haberland (California Polytechnic State University, NumFOCUS)
To develop an open source, unified framework for cell-cell communication modeling and exploration that is available to the entire scientific community.
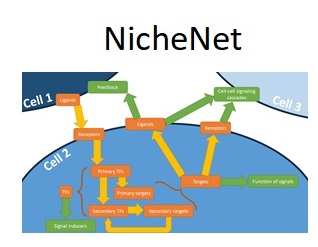
Yvan Saeys (VIB-Ghent University)
To address accessibility gaps in Bokeh, Panel, and Holoviz by incorporating accessibility affordances to ensure everyone can benefit from their powerful data visualization capabilities.

Tania Allard (Quansight Labs)
To make UpSet plots accessible to low-vision and blind users, and to simplify authoring UpSet plots.

Alexander Lex (University of Utah)
To address new challenges posed by replicated single-cell RNA-seq data and by mass spectrometry proteomics.


Gordon Smyth (Walter & Eliza Hall Institute of Medical Research)
To support the onboarding, inclusion, and retention of people from historically marginalized groups on scientific Python projects and structurally improve the community dynamics of NumPy, SciPy, Matplotlib, and pandas.
Melissa Mendonça (Quansight, NumFOCUS)
To advance array interoperability within the PyData Ecosystem by accelerating Array API adoption in key libraries used in biomedical research and building infrastructure for measuring API compliance.

Athan Reines (Quansight)
To support the QIIME 2 user and developer communities by enabling sharing of automatically tested third-party content on the QIIME 2 Library, and hosting our first-ever co-convened user and developer workshop and networking event.

Greg Caporaso (Northern Arizona University)
To enhance the anndataR package for improved interoperability and functionality in handling single-cell data within the R programming environment.
Robrecht Cannoodt (Open Collective Europe Foundation)
To support the growth, sustainability, and diversity of the Apache Arrow project by expanding an apprenticeship program, which recruits developers from underrepresented groups and trains them to be open source software maintainers.

Wes McKinney (Ursa Labs)
To develop software that is able to automatically integrate existing open source software into the Galaxy platform. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Daniel Blankenberg (The Cleveland Clinic)
To develop an automated tool that uses optimization to calibrate models of the musculoskeletal system and improve simulation results, and disseminate the tool to the OpenSim community with documentation, examples, case studies, and outreach events.

Thomas Uchida (University of Ottawa)
To develop a modular and extensible variant caller for microbial genome data that is sequencing technology agnostic and underpinned by an extensive validation and test suite.

Torsten Seemann (The University of Melbourne)
To develop key infrastructure updates and collaboration resources for state-of-the-art Bayesian modeling software libraries.



Christopher Fonnesbeck (NumFOCUS)
To provide ongoing maintenance and community support for the bcbio-nextgen toolkit, focusing on existing variant calling functionality and improving the epigenomic pipelines.

Shannan Ho Sui (Harvard T.H. Chan School of Public Health)
To reengineer the Bioconductor build system for nightly continuous integration, production, and distribution of tarballs and binaries for over 1,700 user-contributed software packages.

Vincent Carey (Brigham and Women's Hospital)
To provide Bioconductor training globally by redeveloping the website and developing infrastructure to deliver high quality community-led training in local languages.

Aedin Culhane (Dana-Farber Cancer Institute, Harvard University)
To increase participation of underrepresented groups in genome data science research through alliances with organizations advancing diversity in science, increased mentoring activities for developers, and enhanced governance of Bioconductor.

Vincent Carey (The Brigham and Women's Hospital)
To extend the open source Bokeh library to cover streaming gridded visualizations for bioscience applications that currently require expensive proprietary tools.

Bryan Van de Ven (NumFOCUS)
This grant supports implementation of biomechanical analysis features in ITK-SNAP, an open source application for medical image segmentation, with the goal of streamlining image processing, anatomical modeling, and tissue mechanics analysis from clinical image data.

Alison Pouch (University of Pennsylvania)
To bridge the gap between the desktop and the web for large-scale GPU scientific visualization by developing a web version of VisPy 2.0 based on Python, WebAssembly, WebGPU, and Datoviz.

Cyrille Rossant (NumFOCUS)
To create a versatile, inclusive, open source microscopy platform for diverse global educational and research communities by modernizing and extending Micro-Manager.
Henry Pinkard (NumFOCUS)
To enhance MNE-Python for clinical neuroscience uses by improving spectral and spectro-temporal data handling, and by providing standardized preprocessing pipelines for data.

Daniel McCloy (University of Washington)
To develop tools, training materials, and mentorship opportunities to help women and nonbinary people in network science to use and contribute code to the igraph open source network analysis library.

Brooke Foucault Welles (Northeastern University)
To enhance metagenomic analysis by integrating cogent3, GraphBin and IQ-TREE to support innovative genomic technologies for monitoring the impact of viral and bacterial diversity on human health.
Gavin Huttley (Australian National University)
To develop extensive functionality, expand user support, and initiate new modes of community outreach for Seurat, an open-source R toolkit for integrative single-cell analysis.
Rahul Satija (New York Genome Center)
To develop extensive functionality, expand user support, and facilitate interoperability for Seurat, an open source R toolkit for integrative single-cell analysis.

Rahul Satija (New York Genome Center)
To reorganize the libSBML and Deviser code bases for better community involvement, spin out part of libSBML as a reusable component for Deviser and other projects, and establish protocols for long-term sustainability of these important resources.


Sarah Keating (University College London)
To develop statistically robust, computationally efficient, and maximally compatible open source software for the design and analyses of multiplexed single-cell sequencing experiments.
Jimmie Ye (University of California, San Francisco)
To effectively support community building efforts across open source ecosystems in molecular sciences, improve contributor pipelines, and seek synergies and collaboration opportunities in this space.
Karmen Condic-Jurkic (Open Molecular Software Foundation)
To maintain BWA and improve the performance and robustness of BWA and its next major version BWA-MEM2.
Heng Li (Dana-Farber Cancer Institute)
To build Cytoscape Explore, a web-based biological network viewer and editor that will make key aspects of the widely used Cytoscape application accessible to new audiences as part of its evolution from a desktop application to a cloud ecosystem.

Dexter Pratt (University of California, San Diego)
To accelerate single-cell biology methods research and empower their developers with foundational probabilistic AI software.

Eli Bingham (Broad Institute of MIT and Harvard)
To interface recently developed computer vision tools with an open-source biomechanical modeling software, which should facilitate the uptake of markerless motion tracking in biomedicine.
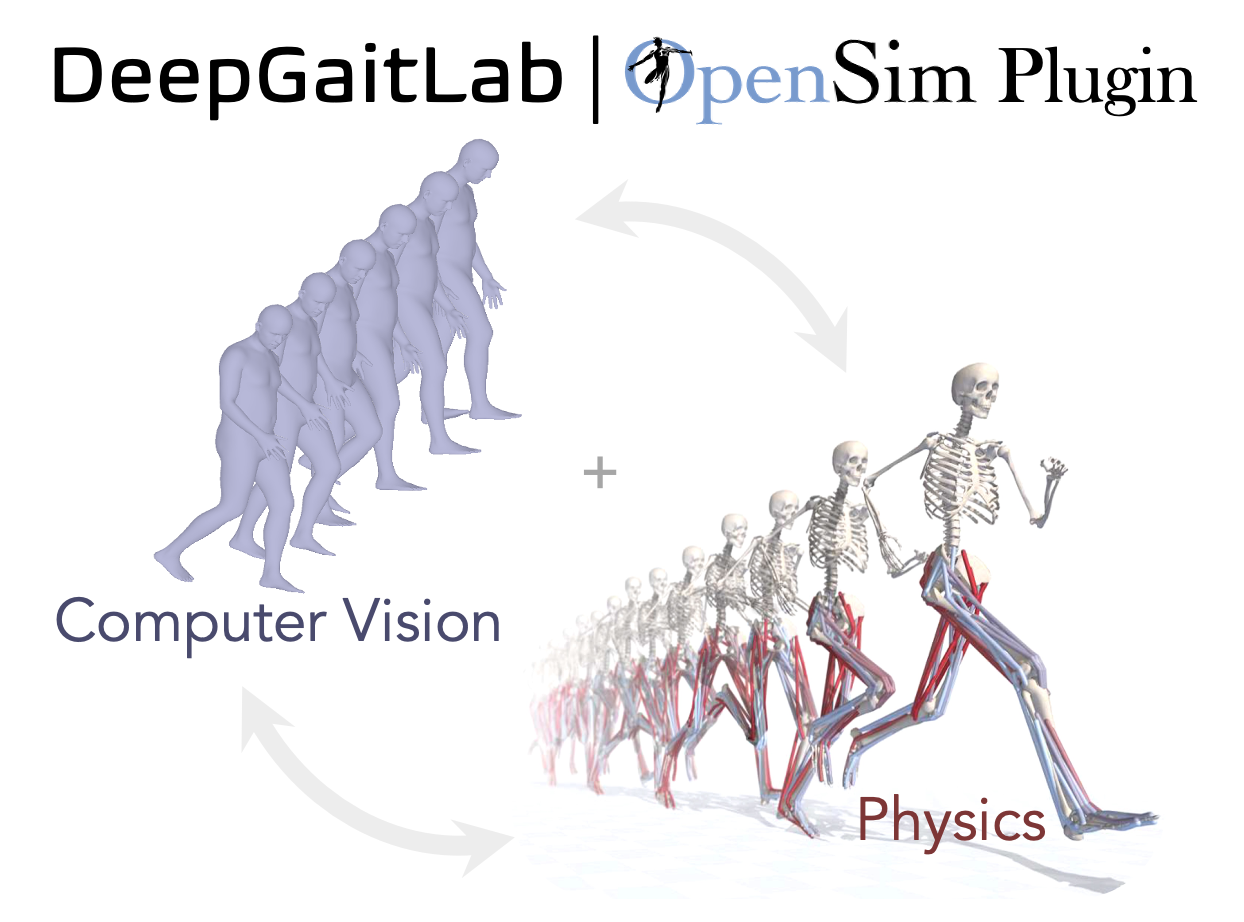
Eni Halilaj (Carnegie Mellon University)
To develop a DeepLabCut AI Residency Program for underrepresented groups in machine learning and computer science in order to recruit, fund, and nurture the next generation of open source leaders.

Mackenzie Mathis & Alexander Mathis (École Polytechnique Fédérale de Lausanne (EPFL))
To provide maintenance, user-focused extensions, education, and support of the growing DeepLabCut software community.

Mackenzie Mathis (École Polytechnique Fédérale de Lausanne (EPFL))
To support the maintenance, new extensions, and education of users of the DeepLabCut software community.

Mackenzie Mathis (Harvard University & Swiss Federal Institute of Technology Lausanne (2020))
To support code maintenance, a new code cookbook, and user education for the DeepLabCut software community and set the foundation towards becoming a sustainable software package for years to come.

Mackenzie Mathis (Harvard University & Swiss Federal Institute of Technology Lausanne)
To expand the global Bioconductor-Carpentries training program, increase equity and accessibility using culturally sensitive AI translation, and build capacity for workshops in Africa

Aedin Culhane (University of Limerick)
To establish DL4MicEverywhere, a containerized deep learning for microscopy toolkit extending the easy-to-use ZeroCostDL4Mic, with interactive notebooks for training and deployment across platforms. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Ricardo Henriques (Instituto Gulbenkian de Ciência)
Single-cell biology is the application of technologies that enable multi-omics investigation at the level of a single cell. This project will streamline trajectory inference from single-cell omics data by improving integration with upstream and downstream analysis pipelines.

Yvan Saeys (Vlaams Instituut voor Biotechnologie)
To enable portability of complex biomedical workflows across different clouds and on-premise environments via better documentation, community support, and tooling for Common Workflow Language (CWL) with examples using Arvados and from the Personal Genome Project.

Sarah Wait Zaranek (Curii Corporation)
To improve ease of use and interoperability of these packages, make methodological responses to new data challenges, refresh the documentation and structure of these packages, and prepare training materials.
Gordon Smyth (Walter and Eliza Hall Institute of Medical Research)
To use QIIME 2 as an on-ramp to scientific computing for Native American students by engaging locally with schools primarily serving Native Americans, while expanding the global QIIME2 user, developer, and educator communities.

Greg Caporaso (Northern Arizona University)
To increase diversity in computational mass spectrometry through teaching and mentoring with the open-source framework OpenMS.

Hannes Rost (University of Toronto)
To enhance Giotto by implementing a novel data structure and framework for the abstract representation and analysis of emerging datasets from multi-modal and multi-resolution spatial technologies.

Ruben Dries (Boston Medical Center Corporation / Boston University)
To provide a series of GPU accelerated routines for signal processing and interpolation in CuPy to be a foundation for the research community.

Kenichi Maehashi (NumFOCUS)
To enhance the Multiscale Object-Oriented Simulation Environment software package through development of new GUIs, data and model import tools, and tutorials for computational modeling in neuroscience and systems biology. Note: This proposal was funded by The Kavli Foundation as part of our co-funded EOSS Cycle 6.
Upinder Bhalla (National Centre for Biological Sciences)
To improve Spyder support for connecting to a remote machine to develop, execute, and debug Python code, as well as installing packages, managing environments and interacting with the remote filesystem.
Carlos Cordoba (Quansight, LLC)
To ensure ongoing maintenance and community growth of Bactopia, focusing on integrating visual reports and comprehensive training materials.

Robert Petit (NumFOCUS)
To develop training materials, perform software maintenance, expand outreach, and provide community support for the Open Health Imaging Foundation (OHIF) web-based medical imaging framework including its underlying libraries (e.g., Cornerstone).


Gordon Harris (Massachusetts General Hospital)
To make significant improvements to the SciML project, which is leveraged by pharmacologists in academia and industry for simulation of virtual clinical trials, drug design, and systems biology modeling.

Samuel Isaacson (Boston University, NumFOCUS)
To enhance bedtools’ functionality, documentation, and access to data, which will empower and expand the user community.


Aaron Quinlan (University of Utah)
To provide significant modernization and enhanced usability of the R packages mixtools and tolerance for improved utilization and accessibility within the biomedical and health research communities.


Derek Young (University of Kentucky Research Foundation)
To extend DESeq2 functions to develop interfaces with Bioconductor’s rich experiment and annotation data, including single-cell datasets and genomic annotations, all leveraging tximeta’s metadata functionality for computational reproducibility.

Michael Love (The University of North Carolina at Chapel Hill)
To support continued maintenance and development of pandas, an open source, BSD-licensed library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language.

Tom Augspurger (NumFOCUS)
To support the growth and health of pandas, the foundational library for tabular data structures in the Scientific Python Ecosystem, by funding continued maintenance and community building efforts.

Joris Van den Bossche (NumFOCUS)
To build a repository of infectious disease models in Stan that will provide researchers with in-depth examples, helping them to draw sound conclusions and make better predictions from their data.

Mitzi Morris (Columbia University)
To enable interactive analysis and exploration of phylogenetic data at the genomics and metagenomics scale.

Jaime Huerta-Cepas (Centro de Biotecnología y Genómica de Plantas (CBGP, UPM/INIA-CSIC))
To support the release and maintenance of a new version of the ETE toolkit including updated documentation and new features such as tree diff, tree-like regular expression searches, and large tree visualization.

Jaime Huerta Cepas (Centro de Biotecnología y Genómica de Plantas)
To make scientific Python documentation more valuable by improving the user experience of linking between projects, and promote this ability within the scientific Python community.

Eric Holscher (Read the Docs, Inc)
To expand on existing web infrastructure by building highly intuitive and responsive data visualization tools for mining ‘shotgun’ metagenomic data.
Daniel Beiting (University of Pennsylvania)
To expand the capabilities and improve the robustness and maintainability of the salmon software ecosystem, further widening its scope of applicability and improving the user and developer experience.

Robert Patro (University of Maryland)
Open mHealth created an open data standard and community for patient-generated data, and the Digital Biomarker Discovery Pipeline will enable transformation of that data into indicators of health outcomes and evaluation of novel digital biomarkers.
Jessilyn Dunn (Duke University)
To extend Galaxy, a web-based computational workbench used by thousands of scientists across the world, so that it can analyze large datasets and connect with other analysis tools.

Jeremy Goecks (Oregon Health & Science University)
To improve the tooling around the conda ecosystem to better serve the millions of users in biological sciences, data sciences, physics, robotics and other scientific disciplines.

Wolf Vollprecht (conda-forge core, QuantStack, NumFOCUS)
To provide an efficient neuroimage analysis pipeline for the medical imaging communities by consolidating advanced DL-methods into a single user-friendly, maintainable, open source software framework.

Martin Reuter (German Center for Neurodegenerative Diseases)
To develop an easy-to-use and validated neuroanatomical framework for the multidimensional image viewer napari, bringing state-of-the-art image analysis plugins to neuroscience.
Adam Tyson (Sainsbury Wellcome Centre & Gatsby Computational Neuroscience Unit, University College London)
To create a consistent, type-annotated, discoverable, and extensible API for scikit-image and facilitate interoperability in the broader image analysis ecosystem.

Juan Nunez-Iglesias (Monash University / NumFOCUS)
To sustain maintenance, improve community support, enhance usability and robustness, and add improvements for the future framework.

Henrik Bengtsson (University of California, San Francisco)
To inspire a diverse global community to collaborate in genome medicine, research, and education by packaging CGAP’s portal and cloud infrastructure as self-serve, orchestrated, open source software.

Dana Vuzman (Harvard Medical School)
To maintain the popular scikit-image Python library for microscopy and medical imaging data and bring significant improvements via development of a backend system enabling multi-threading and GPU acceleration, an improved release process for more rapid cycles, and expanded biomedical examples.

Gregory Lee (Quansight)
To support GPU-accelerated Bioconductor packages through continuous integration, user-friendly packaging of system-level dependencies, and foundational packages for Bioconductor GPU programming.

Levi Waldron (CUNY Graduate School of Public Health and Health Policy)
To enable end-users of ImageJ/Fiji, Icy and napari to process biological imaging time-lapses or large-scale image data tile-by-tile on multiple graphics processing units (GPUs) using CLIJ.

Robert Haase (Cluster of Excellence “Physics of Life”, Technische Universität Dresden)
To expand our community so new individuals can meaningfully contribute code, documentation, workflows and other software artifacts by hiring a dedicated software engineer.

Sarah Wait Zaranek (Curii Corporation)
To optimize GSVA functionality to analyze single-cell and spatial transcriptomic data sets, increasing its robustness and scalability and improving user interface and documentation.
Robert Castelo (Universitat Pompeu Fabra)
To enhance the HTSJDK Java toolkit for genomics with an extensible plugin framework that will enable support for emerging technologies required by contemporary analysis methods, such as long reads, graph/circular references, and epigenetic modifications.
Eric Banks (Broad Institute of MIT and Harvard)
To integrate ilastik with napari and Dask, replacing the outdated internal viewer and task scheduler by modern, community-supported alternatives with the aim to reduce technical debt, engage with the community, and deliver a superior user experience for the bioimage analysis community.

Anna Kreshuk (European Molecular Biology Laboratory)
To enable multi-scale interactive machine learning on large datasets in ilastik through full exploitation of state-of-the-art pyramidal file formats and viewers, and extend functionality to other bioimage analysis tools.

Anna Kreshuk (European Molecular Biology Laboratory)
To make ilastik more interoperable and its results more reusable and reproducible through the development of Python APIs and general improvement of the third-party developer experience.

Anna Kreshuk (European Molecular Biology Laboratory)
To improve Bokeh in key areas that are relevant to bioscience research and to secure a solid foundation for long-term project health and sustainability by engaging in important maintenance and fostering new contributors.

Bryan Van de Ven (Nvidia)
To maintain and improve the three proposed software projects: minimap2, BWA and hifiasm, and extend them to new architectures and new data types.
Heng Li (Dana-Farber Cancer Institute)
To improve OpenRefine to empower users without programming experience to publish research datasets along with verifiable and reproducible workflows, and to automate such workflows.

Antonin Delpeuch (Code for Science & Society)
To facilitate the detection and characterization of pathogens in microbiome data, while supporting community development and dissemination of accessible and reproducible bioinformatics applications.
J. Gregory Caporaso (Northern Arizona University)
Nicholas Bokulich (ETH Zurich)
To scale technical and social support for new analyses in Nilearn including the general linear model, giving access to a broad statistical framework for neuroimagers within the open source Python ecosystem.

Jean-Baptiste Poline (McGill University)
To improve the flexibility and utility of bedtools for large-scale genomic analyses.

Aaron Quinlan (University of Utah)
To improve the robustness and usability of NumPy by continuing to work in documentation and community building, modernizing its integration with Fortran tools via numpy.f2py, and ensuring the sustainability of both NumPy and OpenBLAS.


Melissa Mendonça (Quansight)
To enhance usability of MNE-Python through improvements to its computational efficiency, API, interactive visualization capabilities, and the clarity and consistency of documentation.

Daniel McCloy (University of Washington)
To complete the design, implementation, and rollout of pip's next-generation dependency resolver, and permanently improve pip's maintainer capacity and user experience.
Ernest W. Durbin III (Python Software Foundation)
To improve the user experience of UCSC Xena and better engage users by implementing the redesign of two core features using UX principles, standardizing training materials, and publishing a blog highlighting research use cases.

Jingchun Zhu (University of California, Santa Cruz)
To bring systematically marginalized voices of disabled scientists into scientific computing communities via building and applying accessibility tools, standards, and community contribution practices in the Jupyter ecosystem.

Tania Allard (Quansight LLC)
To improve diversity of computational biology and open source development, providing professional industry opportunities for talent underrepresented within the field of genomics.

Carl Kingsford (Ocean Genomics)
To build a new class of macromolecular modeling methods to study the interplay of structure, dynamics, cellular assemblies, and disease from the subatomic to nanometer scale.
James Fraser (University of California, San Francisco)
To integrate the WebProtégé ontology editor with other open source tools that together constitute an ecosystem that is used widely to develop and manage biomedical ontologies.

Mark Musen (Stanford University)
To develop an open standard and API for phylogenetic models and improve the speed and scalability of the IQ-TREE software for phylogenetic inference from ultra-large genomic data.
Minh Bui (Australian National University)
To broaden participation in the JupyterHub community by establishing a role dedicated to strategy and stewardship for pathways into and throughout the community, as well as programs that provide onboarding and mentorship for historically underrepresented groups.

Chris Holdgraf (NumFOCUS)
To improve community support and technical maintenance across the JupyterHub repositories.


Chris Holdgraf (University of California, Berkeley; NumFOCUS)
to increase usability of LinkML (Linked data Modeling Language), an open, extensible framework for modeling, validating, and distributing data that is reusable and interoperable. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.
Sierra Moxon (Lawrence Berkeley National Laboratory)
To maintain the established infrastructure and optimize the current features of the popular peak caller MACS for gene regulation studies, while focusing on building the data structure and features for single-cell data analysis.

Tao Liu (Roswell Park Alliance Foundation)
To enhance the infrastructure to support the continuous development and growing community of the popular algorithm MACS for gene regulation studies, in order to expand its features and adapt to new technologies such as single-cell ATAC-seq.

Tao Liu (Roswell Park Alliance Foundation)
To put Rocker, the de facto standard for reproducible, containerized R analyses, on a path to sustainable maintenance through refactoring, improving the quality of documentation, expanding the community, and targeting new hardware platforms.

Carl Boettiger (University of California, Berkeley)
To further the sustainability and usability of scikit-learn by reducing the maintenance backlog and extending its machine learning models and pipelines to support more complex datasets.

Thomas Fan (Quansight)
To improve the bcbio-nextgen toolkit, focusing on maintaining existing variant calling functionality and extending support for structural and RNA-seq variant analyses.

Rory Kirchner (Harvard Chan School of Public Health)
To enable Matplotlib to continue as the core plotting library of the scientific Python ecosystem by addressing the maintenance backlog and planning Matplotlib's evolution to meet the community’s visualization challenges for the next decade.

Thomas A. Caswell (Brookhaven National Laboratory, NumFOCUS)
To enable Matplotlib to continue as the core plotting library of the scientific Python ecosystem for researchers in biomedical imaging, microscopy, and genomics by addressing the maintenance backlog and beginning Matplotlib's evolution to meet the community’s visualization challenges for the next decade.

Thomas A. Caswell (Brookhaven National Laboratory, NumFOCUS)
To support the continued maintenance, growth, development, and community engagement of Matplotlib, the foundational plotting library of the Scientific Python Ecosystem.

Thomas Caswell (Brookhaven National Laboratory, NumFOCUS)
To improve and maintain high-performance tools for analysis of biological systems at the molecular scale and incentivize scientists to drive transparent and reproducible research.

Oliver Beckstein (Arizona State University, NumFOCUS)
To grow the MDAnalysis community sustainably.

Richard Gowers (Open Molecular Software Foundation)
to develop the first user-friendly, open source software that synthesizes and evaluates the evidence about the effectiveness and safety of medical interventions and informs treatment decision-making. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.
Alexander Sutton (University of Leicester)
To support the open source µManager optical microscopy acquisition platform to improve its architecture, infrastructure, and support to ensure many years of growth, both in user base and capabilities.

Kevin Eliceiri (University of Wisconsin, Madison)
To construct a solid foundation for the next generation of Protege using a modern web stack that will make Protege easier to maintain, extend, and — crucially — make it easier for third parties to contribute to the code base.

Mark Musen (Stanford University)
To modernize the igraph interfaces to make network analysis easier.

Vincent Traag (Leiden University)
To keep ChimeraX molecular and microscopy analysis software current with the latest technology and facilitate the migration of tens of thousands of Chimera users to ChimeraX.

John Morris (University of California, San Francisco)
To provide open-source, interoperable, and extensible statistical software for quantitative mass spectrometry, which enables experimentalists and developers of statistical methods to rapidly respond to changes in the evolving biotechnological landscape.

Olga Vitek (Northeastern University)
To develop features for an improved user and developer experience in the Neurodesk platform allowing for easier, portable, scalable, and reproducible neuroscience and biomedical imaging analysis. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.
Steffen Bollmann (The University of Queensland)
To support the Bio-Formats user community and develop new formats to make proprietary file formats obsolete.
Jason Swedlow (University of Dundee)
To enable the analysis of thousands of next generation data-independent acquisition (DIA) mass spectrometry measurements by implementing algorithms, visualization tools, and cloud containers based on OpenMS and the OpenSWATH algorithm.

Hannes Rost (University of Toronto)
To enhance the DeepLabCut ecosystem with a new 3D pose module, integration with generative AI (AmadeusGPT), providing codebase and DLC-Live! code upkeep, and continuing our DEI AI Residency Program. Note: This proposal was funded by The Kavli Foundation as part of our co-funded EOSS Cycle 6.

Mackenzie Mathis (Swiss Federal Institute of Technology, Lausanne)
To develop and integrate open computational geometry and simulation technology enabling digital twins in biomedicine – next-generation imaging-based modeling, simulation, optimization, and learning. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Marie E. Rognes (Simula Research Laboratory)
To keep the current momentum of initiatives and push forward with new actions for accessibility, internationalization, mentorships and ambassadors.


Ellen Sherwood (KTH Royal Institute of Technology)
To support a fast-growing, community-building software for infrastructure agnostic, open source biomedical pipelines.


Ellen Sherwood (KTH Royal Institute of Technology)
To continue support for a fast-growing community, building open source software for infrastructure agnostic, open source biomedical analysis workflows.


Ellen Sherwood (KTH Royal Institute of Technology)
To coordinate and foster next-generation file formats while increasing community access to public imaging data.

Josh Moore (German BioImaging Society)
To solidify NiPreps by boosting community growth, securing maintenance, and developing new components to expand the diversity of supported data such as imaging parameters, modalities, populations, and species.
Oscar Esteban (Lausanne University Hospital and University of Lausanne)
To develop a general-purpose Java library for the community standard image file format OME-Zarr, and robustly integrate it into the widespread image processing and analysis software Fiji.

Pavel Tomancak (Max Planck Institute of Molecular Cell Biology and Genetics)
To enhance discoverability and use of Bioconductor packages and data and teaching resources by tagging them with formal vocabulary and concept relations in ontologies like EDAM.

Vincent Carey (Harvard Medical School)
To partner with hack.diversity to serve as curriculum designers and mentors, equipping their Fellows to contribute to open source web-based medical imaging, as well as mentor existing global OHIF contributors.


Gordon Harris (Massachusetts General Hospital)
To develop, upgrade, and migrate documentation; perform software maintenance; and provide community support for the Open Health Imaging Foundation web-based medical imaging framework.

Gordon Harris (Massachusetts General Hospital)
This project will improve accessibility, interoperability, efficiency, and sustainability of the biomedical image registration software elastix by providing complete support of Python, better integration with other software, code improvements, and a focus on community.

Marius Staring (Leiden University Medical Center)
To improve accessibility, interoperability, efficiency, and sustainability of the biomedical image registration software elastix, by making it a library-first package, allowing integration with other software and improving its performance.

Marius Staring (Leiden University Medical Center)
To improve accessibility, efficiency, and sustainability of the biomedical image registration software Elastix, by improving interaction possibilities, speed, and accessibility via a web browser. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Marius Staring (Leiden University Medical Center)
To develop open medical ontologies and analytics that enable large-scale generation of real-world evidence on disease and the effects of medical interventions across the world’s electronic health data.

George Hripcsak (Columbia University)
To support development, outreach, and user support for the kallisto RNA-seq and single-cell RNA-seq software project.
Lior Pachter (California Institute of Technology)
To create a sustainable open-source community resource for deep annotation of genetic variants.

Rachel Karchin (Johns Hopkins University)
To administer and expand the scope of the Tidyomics Project, enabling computational biologists to employ a user-friendly grammar to manipulate rich biological datasets across omics and platforms. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Michael Love (University of North Carolina Chapel Hill)
To grow the OpenMM development community and expand the OpenMM software ecosystem to better serve user needs. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.
Thomas Markland (Stanford University)
This team will support the continued development of OpenMM to better serve its broad biomolecular modeling community, as well as support its extension to integrate machine learning that will enable genomic-scale biomolecular modeling, simulation, and prediction.
Thomas Markland (Stanford University)
To continue to diversify contributors by building capacity in project management, as well as offering internships and eliminating cultural or linguistic biases in the Open Refine tool.

Antonin Delpeuch (Code for Science & Society)
To develop accurate, fast, and researcher-friendly open tools for creating and simulating neuromuscular and musculoskeletal models to address biomedical questions in human and animal mobility.

Ajay Seth and Adam Kewley (Delft University of Technology)
To improve the usability, computational performance, maintenance and outreach of the open source software OpenSim and to support the education and training of its users around the world.

Ajay Seth (Delft University of Technology)
To hire an outreach coordinator and (part of) a software developer for the Apollo genome annotation editor, provide developer- and user-oriented workshops and training, develop a plugin framework, and integrate protein visualizations.
Ian Holmes (University of California, Berkeley)
To upgrade the interactive documentation experience of IPython and Jupyter to allow inline graphs, navigation, and indexing, and to support features currently only available on hosted websites.

Matthias Bussonnier (NumFOCUS, Quansight Labs)
To update NetworkX algorithms, increase performance of key biomedically related tools such as community detection (CD) and subgraph-isomorphism (SI), and develop a roadmap for improved visualization. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Daniel Schult (Colgate University)
To develop an open Python library and community between spectroscopy and fluorescence microscopy users that is both accessible and self-sustainable in the long term.

Leonel Malacrida (Institut Pasteur Montevideo)
To develop an end-to-end, predictive computational ecosystem for quantitative spatiotemporal modeling of spatial and single-cell multiomics.
Jonathan Weissman (Whitehead Institute for Biomedical Research)
To sustain the scikit-learn library through maintenance, improvement, and extension, notably in the domain of predictive model evaluation and inspection. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.

Guillaume Lemaitre (Inria Foundation)
To support the maintenance, development and dissemination of ImgLib2 and BigDataViewer, key infrastructural software components for visualization and analysis of large image data on the Java-based platforms Fiji, KNIME, and Icy.


Pavel Tomancak (Central European Institute of Technology)
This project will solidify the use and future development of the cross-language igraph package to support network analysis in all scientific domains, including biomedical and life sciences.

Vincent Traag (Leiden University)
To create a smoother experience for PsychoPy users by detecting potential problems in user-created experiments before they are launched, and by increasing the test coverage within the application code itself.
Jonathan Peirce (University of Nottingham)
To enable researchers to more deeply interrogate complex biomedical images by improving the extensibility, robustness, and interoperability of QuPath.

Peter Bankhead (University of Edinburgh)
To accelerate biomedical research, biomarker discovery, and the translation of artificial intelligence into clinical practice by enhancing the QuPath open source platform and by integrating it with other CZI-funded software.
Peter Bankhead (University of Edinburgh)
To hire a dedicated Communities Champion, running code-sprints and contributor workshops and translating our desktop app into additional languages.

Jonathan Peirce (University of Nottingham)
This team will build a real-time data model for Jupyter to lay the foundation for real-time collaboration on notebooks.

Saul Shanabrook (Quansight)
To significantly speed up IQ-TREE to enable real-time genomic epidemiology during ongoing outbreaks such as COVID-19, and to introduce continuous integration and a testing framework to ease software maintenance for all developers.
Minh Bui (Australian National University)
To attract new users and contributors to the VisPy project through software improvements, community outreach, and instructional materials.

David Hoese (University of Wisconsin - Madison)
To establish teaching material, improve documentation, and minimize maintenance effort of the Bioconda project by extending automation of code review, testing, and building.

Johannes Köster (University of Duisburg-Essen, Bioconda Core Team)
To meet the needs of the scientific community over the next decade, this team will revitalize NetworkX — the fundamental network analysis tool in Python — by growing its developer community, refactoring code, improving performance, and making a major release.

Stefan van der Walt (University of California, Berkeley)
To advance support and development of the open source Salmon and Alevin software for gene expression quantification of single-cell and bulk RNA-seq.

Carl Kingsford (Ocean Genomics)
To establish Zarr as a foundation for scientific data storage, with clear data format and protocol specifications, implementations in multiple programming languages, and a community process for evolving to support new scientific applications.

Ryan Williams (Mount Sinai School of Medicine)
To refactor Orange Data Mining toolbox to include the latest Python libraries for parallel, server-based data analysis, allowing it to scale to large biomedical datasets.
Blaž Zupan (University of Ljubljana)
To attract new contributors by improving OpenRefine's documentation, and implement a new data model to improve the scalability, transparency, and reproducibility of OpenRefine workflows.

Antonin Delpeuch (Code for Science and Society)
To provide dedicated community support and maintenance for the Dask project and support growth in the biological sciences field.

Matthew Rocklin (NumFOCUS)
To scale the core scverse data structures (AnnData, MuData, and SpatialData) for out-of-core/cloud-based analysis to accelerate everything from common analysis tasks to foundational model training. Note: This proposal was funded by Wellcome Trust as part of our co-funded EOSS Cycle 6.
Fabian Theis (Helmholtz Munich)
To expand Scanpy’s core infrastructure and community platforms for stability, versatility, and sharing knowledge.
Fabian Theis (Helmholtz Zentrum Munich)
To improve the open-source machine learning library scikit-learn and aid in maintaining the project, while considering the new implementation of Gradient boosting.

Andreas Mueller (Columbia University)
To better serve biomedical applications, SciPy will add important new features, perform essential maintenance, and disseminate the work to biomedical researchers and software developers.

Matt Haberland (California Polytechnic State University, San Luis Obispo)
To maintain and further develop a community resource for probabilistic analysis of single-cell omics data, including an application interface for rapid development of new probabilistic models.

Nir Yosef (University of California, Berkeley)
To enhance Snakemake accessibility with open source tools and engage the community to reduce ad-hoc unstructured code and promote a culture of sustainability in data analysis beyond reproducibility.

Ivan Molineris (University of Turin)
To turn SPAdes and QUAST codebases into scalable, modular, extensible and user-friendly frameworks that will streamline future research and development in genome assembly, analysis and quality assessment.

Anton Korobeynikov (Saint Petersburg State University)
To improve sparse structures in SciPy so they support array semantics, to deprecate SciPy’s sparse metrics, and to assist with sparse array adoption in downstream ecosystem packages.

Stéfan van der Walt (University of California, Berkeley)
To strengthen the social and code foundations of the Nibabel library by extending the API and input/output to better support metadata, supporting outputs from image registration, and through educational outreach.
Matthew Brett (NumFOCUS)
To extend DIPY’s registration framework for generic use in biomedical research, add parallel computing, strengthen maintenance, expand documentation, and improve educational capabilities.

Serge Koudoro (Indiana University - DIPY)
To grow the maturity of the NumPy project through governance, documentation, and website work by improving the robustness of its links with OpenBLAS, and through diversifying the core team beyond the developer role.
Ralf Gommers (Quansight, NumFOCUS)
To champion Bioconductor developers through better community and technical infrastructure, accessibility of documentation, support for developer working groups, and cross-community collaborations.

Maria Doyle (University of Limerick)
To improve Monocle with algorithms, statistical methods, and web-based visualization tools that will enable biologists using single-cell genomics to extract and disseminate new insights from their experiments.
Cole Trapnell (University of Washington)
To accelerate the adoption of Xarray in the biomedical research community.

Joe Hamman (Earthmover PBC)
To expand and strengthen the NetworkX developer community, reinforce connections with the scientific Python ecosystem, improve documentation and training materials for users, and refine development infrastructure and process.

Daniel Schult (NumFOCUS)
To enable the biomedical community to more easily use large-scale computing to efficiently run complex workflows via Parsl.

Kyle Chard (University of Chicago)
To improve the SymPy Python symbolic mathematics library in the key areas of performance, code generation, and documentation.

Aaron Meurer (Quansight)
To develop GATK methods for variant discovery and evaluation in bacteria, resolve inconsistent and diverse results from different research groups, and allow for the sharing of data and analysis globally to control bacterial transmission and antibiotic resistance.

Bhanu Gandham (Broad Institute of MIT and Harvard)
To ensure the continued health of the pandas library by dedicating resources specifically to maintenance and implementing consistent missing data handling for all data types.

Tom Augspurger (NumFOCUS)
To improve Percolator, the dominant software for analyzing spectrum identifications produced by protein tandem mass spectrometry, by making the software faster, more robust, and applicable to more types of mass spectrometry data.

William Noble (University of Washington)
To improve conda-forge and bioconda’s sustainability and transparency by adopting vendor-agnostic and secure infrastructure practices and develop comprehensive maintenance metrics and dashboards.

Jaime Rodríguez-Guerra (Quansight)
To cultivate the next generation of biomedical open source software data scientists through recruitment, mentorship, and training of underrepresented students.

Jingchun Zhu (University of California, Santa Cruz)
To improve the interoperability, efficiency, and robustness of popular bulk and single-cell transcriptomics analysis tools, and to add important new capabilities to the open source ecosystem.

Robert Patro (University of Maryland)
To support biomedical research in low-resource settings with open tools for extracting, normalizing, and de-identifying patient-level clinical data from widely adopted OpenMRS Electronic Medical Records.

Jan Flowers (University of Washington)
To support and further develop a library for high-performance scientific visualization in Python by maintaining the VisPy package and improving documentation within the community.

Kenneth Harris (University College London)
To maintain and enhance seqr, a high quality rare disease genomic analysis platform, for usage across the global scientific community, enabling collaboration for rare disease diagnosis and gene discovery.

Heidi Rehm (Massachusetts General Hospital and Broad Institute of MIT and Harvard)
To grow the use of Xarray in the biosciences as a foundational data model and computational toolkit for multidimensional labeled arrays.

Joseph Hamman (NumFOCUS)
To establish Zarr as a common, cross-community mechanism for storing collections of annotated tensors with consistent access for both local and large-scale cloud data.

Josh Moore (German BioImaging)
Sorry, there are currently no results that match those criteria. Please try clearing all search terms.



